
The discovery of genetic diversity in Ae. tauschii associated with traits of interest can help accelerate wheat improvement. To address this aim, we seek to comprehensively characterize the genetic diversity of this wild relative of wheat.
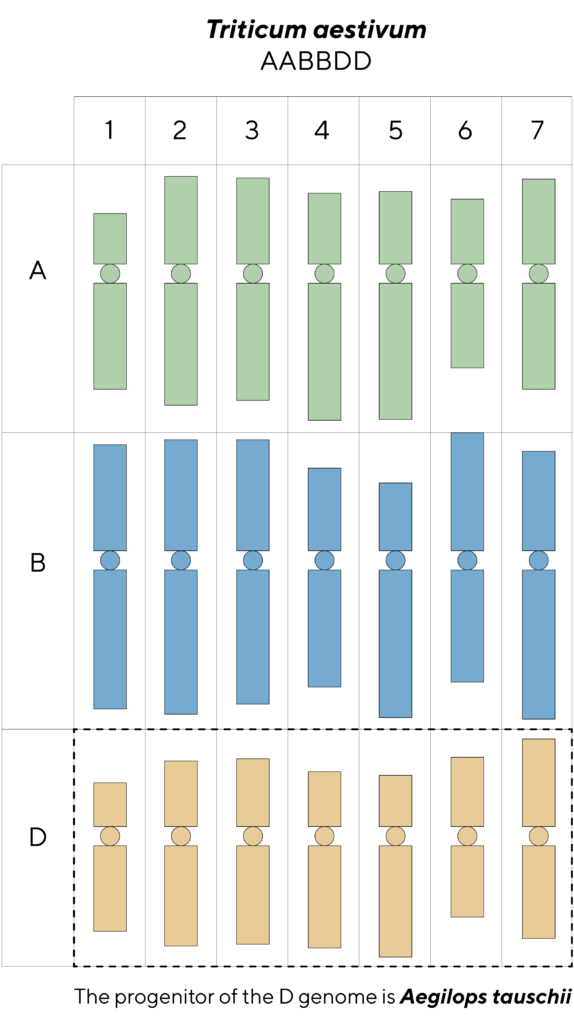
Hexaploid bread wheat, Triticum aestivum (genome constitution AABBDD), emerged from two successive hybridization events from diploid progenitors 5.5 million and 8,000 years ago, respectively.
The progenitor of the D genome is the wild wheat species Aegilops tauschii. This goatgrass has a natural distribution ranging from East Turkey to China and West Pakistan.
Ae. tauschii is classified into three lineages, namely lineage 1 (L1), lineage 2 (L2), and lineage 3 (L3). The wheat D sub-genome resulted from at least two hybridization events with predominantly L2 and L3 donors, respectively.
This wild wheat species is a suitable model system for studying trait variation transferrable to wheat. It has a diploid genome of 4.36 Gb, nearly four times smaller than that of hexaploid wheat (16 Gb).

The Open Wild Wheat Consortium phase 1 was completed in 2021 with the publication of our research article in Nature Biotechnology. The publication took five years in the making and gathered 78 co-authors from 38 institutions around the world.
We established a diverse panel of 242 inbred and genotypically non-redundant accessions of Aegilops tauschii from across its natural geographic range.
The Ae. tauschii panel includes 116 accessions from Lineage 1, 121 from Lineage 2, and 5 from Lineage 3, displaying extensive variation in morphological traits and disease resistance.
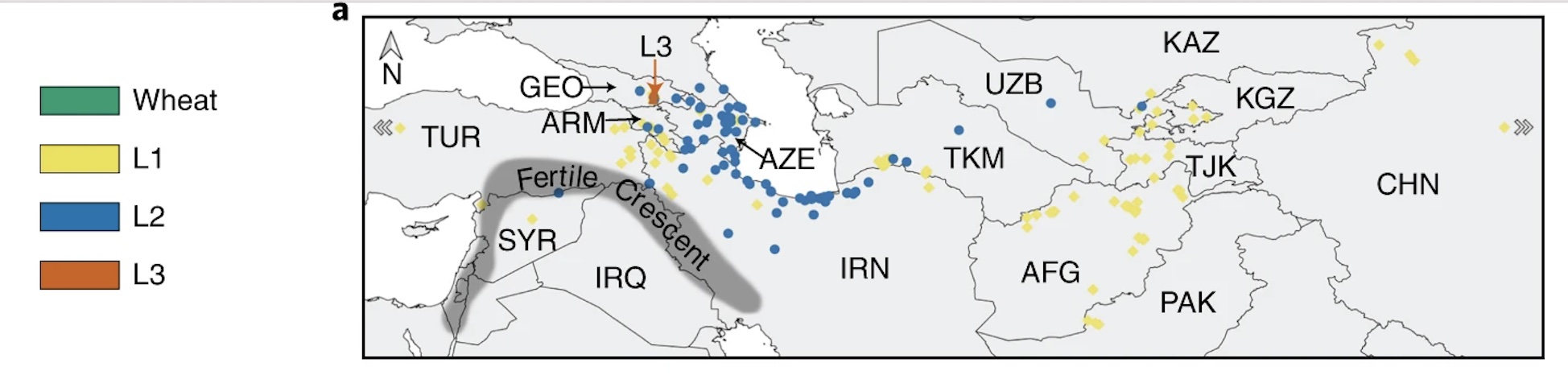
Geographical distribution of the Aegilops tauschii accessions included in the diversity panel. The accessions are color coded according to their lineage. Figure 1a taken from Gaurav et al., 2022.
GERMPLASM
Seed for the OWWC diversity panel is publicly available in the Germplasm Resources Unit (GRU) of the John Innes Centre, UK.
– GRU Collection: Open Wild Wheat Consortium Ae. tauschii Diversity Panel
PUBLICATIONS
Gaurav et al. (2022) Population genomic analysis of Aegilops tauschii identifies targets for bread wheat improvement. Nature Biotechnology.
Delorean et al. (2021) High molecular weight glutenin gene diversity in Aegilops tauschii demonstrates unique origin of superior wheat quality. Communications Biology. 4, 12422
DATA AND CODE
Data generated by the OWWC and published in Gaurav et al. 2022 is available in public repositories as described here.
Scripts and pipelines for the bioinformatics analyses described in Gaurav et al. 2022 are available in the Github repository: wheatgenetics/owwc.
PRESENTATIONS, PRESS RELEASES, AND MEDIA
How bread wheat got its gluten: Tracing the impact of a long-lost relative on modern bread wheat – Press release by JIC Communications (Nov 2021)
2020 BGRI Virtual Technical Workshop:
Exploiting diversity in the bread wheat D-genome progenitor. Presented by Kumar Gaurav. (Oct 2020)
New genomic tool searches wheat’s wild past to improve crops of the future –Press release by JIC Communications (Apr 2018)
The Open Wild Wheat Consortium phase 2 began in January of 2022 and was completed in August of 2024 with the publication of our research article in Nature. Seventy-one co-authors from 31 institutions worldwide collaborated on this study.
The hallmark of the OWWC Phase 2 is the creation of an Aegilops tauschii pangenome resource and the expansion of the diversity panel to 493 genetically non-redundant accessions.


Bread shaped human civilization. It provided the delicious doughy calories that energised the populations of some of the first ancient cities. Ten thousand years later this iconic crop helps to sustain a global population of eight billion.
A remarkable new study has provided compelling evidence as to how it was made possible, revealing the secret that made bread wheat one of the most successful crops on the planet while many others have disappeared or remained in local obscurity.
“Our findings shed new light on an iconic event in our civilisation that created a new kind of agriculture and allowed humans to settle down and form societies,” said Professor Brande Wulff, a wheat researcher at KAUST (King Abdullah University of Science and Technology) and one of the lead authors of the study which appears in Nature.
The secret, according to research by the Open Wild Wheat Consortium (OWWC), lies in the genetic diversity of a wild grass called Aegilops tauschii.
This inconspicuous weed provided bread wheat’s D-genome when it crossed with early cultivated pasta wheat in the Fertile Crescent sometime between eight and eleven thousand years ago.
The chance hybridisation on the banks of the southern Caspian Sea spawned an agricultural revolution as farmers enthusiastically adopted this dynamic new crop with its high gluten content that creates an airier elasticated breadmaking dough.
Within a few hundred, maybe a few thousand years its cultivation was dispersed across a wide new climatic and agricultural range.
This rapid geographical advance has puzzled wheat researchers. There is no wild bread wheat: and the kind of hybridisation event that added the new D genome to wheat’s existing A and B genomes created a genetic bottleneck, caused by a new species contained within a small population and lacking in genetic diversity compared to wild grasses.
This bottleneck effect coupled with the fact that wheat is an in-breeding species – meaning it is self-pollinating – would suggest that bread wheat might struggle outside its Fertile Crescent origins. So how did it become well-travelled?
In solving this conundrum, the international collaboration assembled a diversity panel of 493 unique accessions spanning the geographical range of Aegilops tauschii from north-western Turkey to eastern China.
From this panel the researchers selected 46 accessions reflecting the species traits and genetic diversity, to create a pan-genome, a high-quality genetic map of Aegilops tauschii.
Using this map, they scanned 80,000 bread wheat landraces – locally adapted varieties – held by the public-good wheat breeding institution CIMMYT and collected from around the world.
This data showed that only around 75% of the bread wheat D-genome is derived from the lineage (L2) of Aegilops tauschii which originates from the southern Caspian Sea. The remaining 25% of its genetic make-up is derived from lineages across its range.
“This 25% influx of genetic material from other lineages of tauschii has contributed and defined the success of bread wheat,” said Professor Simon Krattinger, lead author of the study.
“Without the genetic viability that this diversity brings, we would most likely not eat bread on the scale we do today. Otherwise, bread wheat today would be a regional crop – important to the Middle East but I doubt that it would have become globally dominant without this plasticity that enabled bread wheat to adapt.”
A previous study by OWWC revealed the existence of a distinct lineage of Aegilops tauschii geographically restricted to present day Georgia in the Caucasus region – 500 kilometers from the Fertile Crescent.
This Aegilops tauschii lineage (L3) is significant because it has provided bread wheat with the best-known gene for dough quality.
In this study the researchers hypothesised that if this were an historic introgression, akin to a Neanderthal genetic footprint in the human genome, they would find landraces in the CIMMYT collections that had a higher proportion of it.
Data analysis showed that CIMMYT wheat landraces collected from the Georgian region contained 7% L3 introgressions in the genome, seven times more than that of bread wheat landraces collected from the Fertile Crescent.
“We used the L3 tauschii accessions as a guinea pig to track and trace the hybridizations using 80,000 bread wheat landraces,” said Professor Krattinger.
“The data beautifully supports a picture where bread wheat emerges in the southern Caspian, then with migration and agricultural expansion it reached Georgia and here with gene flow and hybridisations with the peculiar, genetically distinct and geographically restricted L3 accessions it resulted in the influx of new genetic material.”
“This is one of the novel aspects of our study and it confirms that using our new resources we can trace the dynamics of these introgressions in bread wheat.”
In addition to solving this age-old biological mystery the new Aegilops tauschii open source pan-genome and germplasm made available by the OWWC, are being used by researchers and breeders worldwide to discover new disease resistance genes that will protect wheat crops against age-old agricultural plagues like wheat rust. They can also mine this wild grass species for climate resilient genes which can be bred into elite wheat cultivars.
“This has been a wonderful collaborative effort by the OWWC, an international cross institute enterprise that has produced the best sequenced resource of a wild wheat relative in the world,” said Professor Wulff.
“Wild relatives such as Aegilops tauschii offer amazing genetic diversity that breeders can exploit with the tools we have now created and which have not been exploited in landraces, there is so much more potential that is untapped.” he concluded.
Origin and evolution of the bread wheat D genome appears in Nature.